Introduction
When the data model of proteomics is relatively complex, such as the protein matrix data of multiple organs and tissues being analyzed together, the protein levels and types of various tissues or organs are quite different, and the conventional bioinformatics methods cannot effectively and efficiently identify the key proteins of each tissue. Here we can use the relatively new analysis methods in genomics, such as weighted gene co expression analysis, unsupervised deconvolution method, to solve this kind of analysis problem.
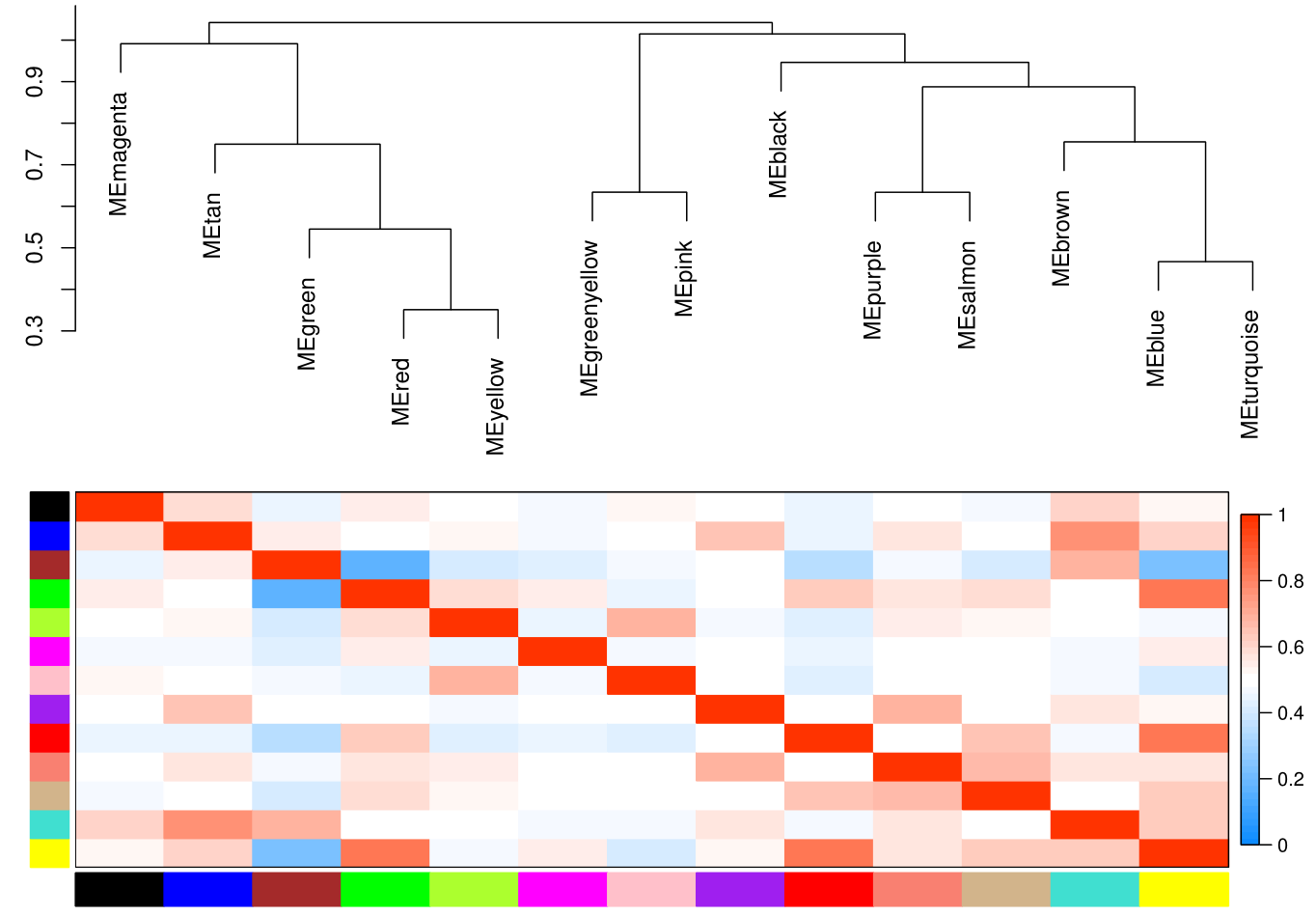
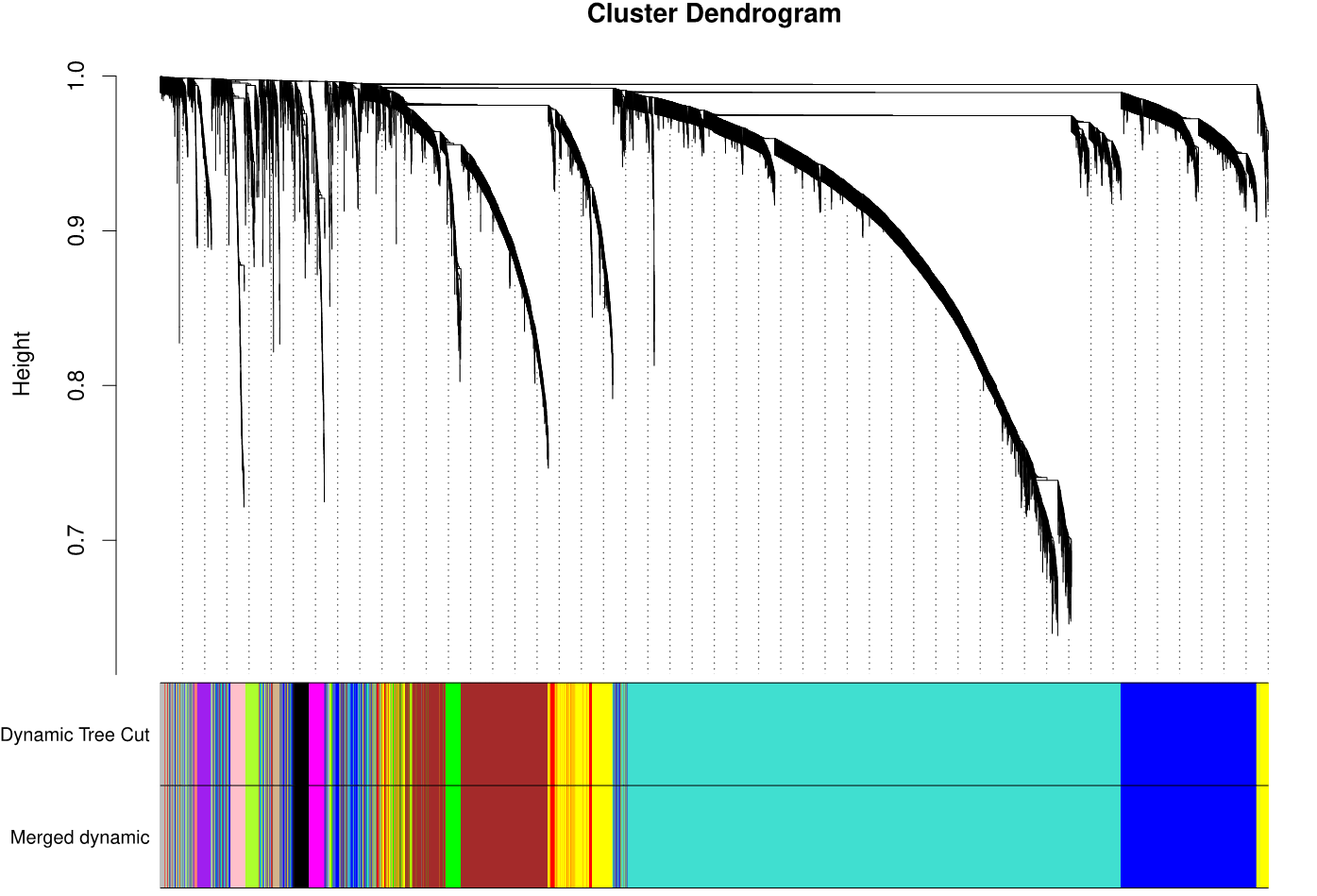
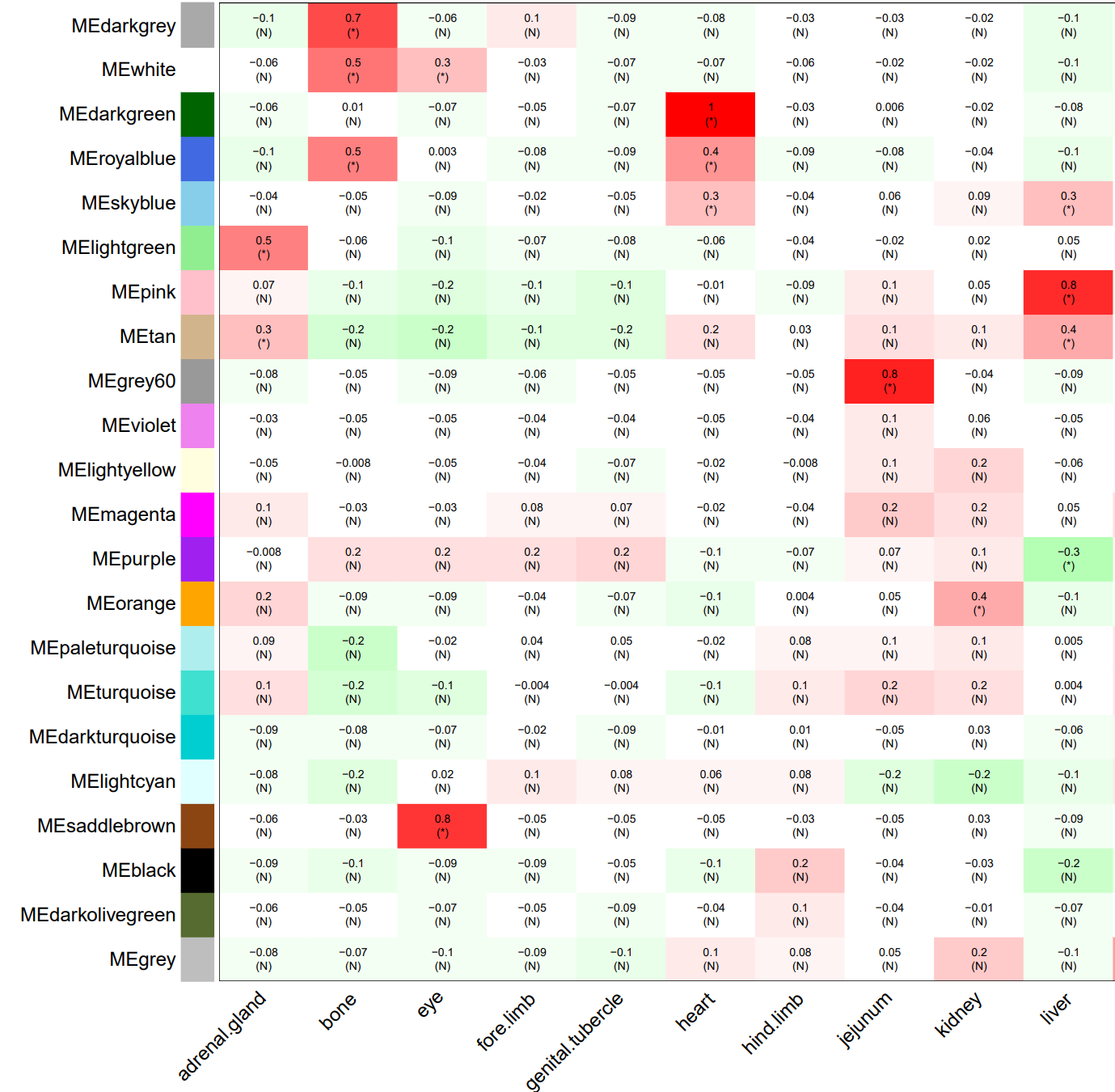
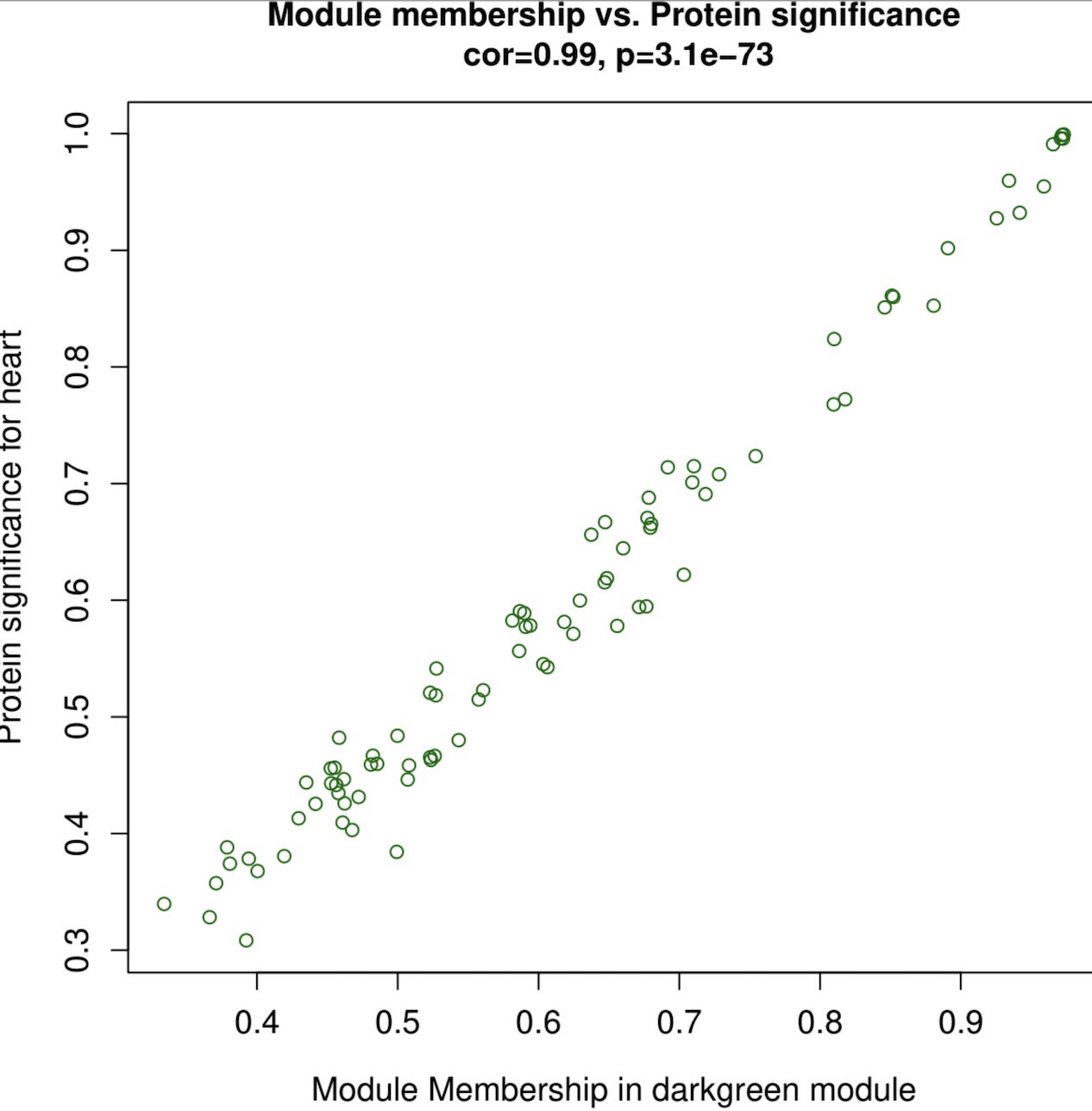
The weighted gene co-expression network analysis (WGCNA) method, assuming that all kinds of proteins in the organism obey the basic assumption of scale-free network topology, is used to search for proteins or gene modules (A, B) that are co expressed based on the correlation algorithm, and explore the relationship between each module and each organ or organization (C), as well as the core protein or gene (D) in each module.
Case